Epigenetic Analysis of the Aging Brain (Part 2)
Single-cell Transcriptomic and Epigenetic Assays Detail the Impact of Aging Throughout the Mouse Brain: A new study employing transcriptomic and epigenetic assays reveals the significant impact of aging in single cells isolated from distinct regions of the mouse brain.
Exploring the Epigenetic Alterations Accompanying Mouse Brain Aging at the Single-cell Level
Normal aging in the brain occurs alongside widespread cellular and molecular changes that contribute to cognitive decline, reduced neural plasticity, and increased vulnerability to neurodegenerative diseases (Mattson and Arumugam). While single-cell transcriptomic profiles have provided some insight (Almanzar et al. and Jin et al.), we require a deeper appreciation of how epigenomic landscapes shift with age across different brain regions and cell types to understand the transcriptional regulatory mechanisms that induce detrimental age-related changes in the brain. Changes in chromatin accessibility, DNA methylation, and histone modification profiles can induce the progressive loss of transcriptional fidelity and cellular identity (Patrick et al.) and, as such, epigenetic alterations represent crucial contributors to the aging process (Wang et al.). Conventional "bulk" epigenetic analyses have hindered cell-type- and region-specific analyses, limiting the specific analysis of, for example, dopaminergic neurons in the substantia nigra in Parkinson's disease, oligodendrocytes in the white matter in multiple sclerosis, and hippocampal neurons in Alzheimer's disease. Can single-cell epigenetic analyses help to capture the precise regulatory mechanisms driving age-related and cell-type- and region-specific shifts in gene transcription in the brain?
To fill this knowledge gap, researchers led by Margarita Behrens (Salk) and Bing Ren (University of California San Diego) recently sought to create a comprehensive single-cell atlas of chromatin accessibility and transcriptional changes during mouse brain aging by integrating scATAC-seq, scRNA-seq, and spatial transcriptomics datasets generated from distinct brain regions in mice at 2, 9, and 18 months of age. Their new study, reported in Cell Reports, now demonstrates how resolving epigenomic changes at single-cell resolution uncovered the regulatory mechanisms underlying aging-associated transcriptional shifts and cellular dysfunction (Amaral et al.). Overall, they reveal age-related loss of master transcription factor binding, stress-induced reprogramming, and heterochromatin instability in glial and neuronal cell types across distinct regions of the mouse brain.
Paired-Tag technology from Epigenome Technologies generates joint epigenetic and transcriptomic profiles at single-cell resolution and detects histone modifications and RNA transcripts in nuclei with efficiency comparable to single-nucleus RNA-seq/ChIP-seq assays. Could integrating Paired-Tag into the experimental scheme employed have provided even deeper epigenetic insights into brain aging at the single-cell level?
Epigenetic Analyses Link Aging to the Loss of Brain Cell Fate
The authors first noted a marked decline in populations of progenitor cells involved in neurogenesis and myelination, as well as widespread, concordant age-associated changes in transcription and chromatin accessibility across neuronal and glial cell types. Genes involved in oxidative phosphorylation and heterochromatin maintenance (including histone methyltransferases and chromatin remodelers) displayed the most consistent downregulation across cell types.
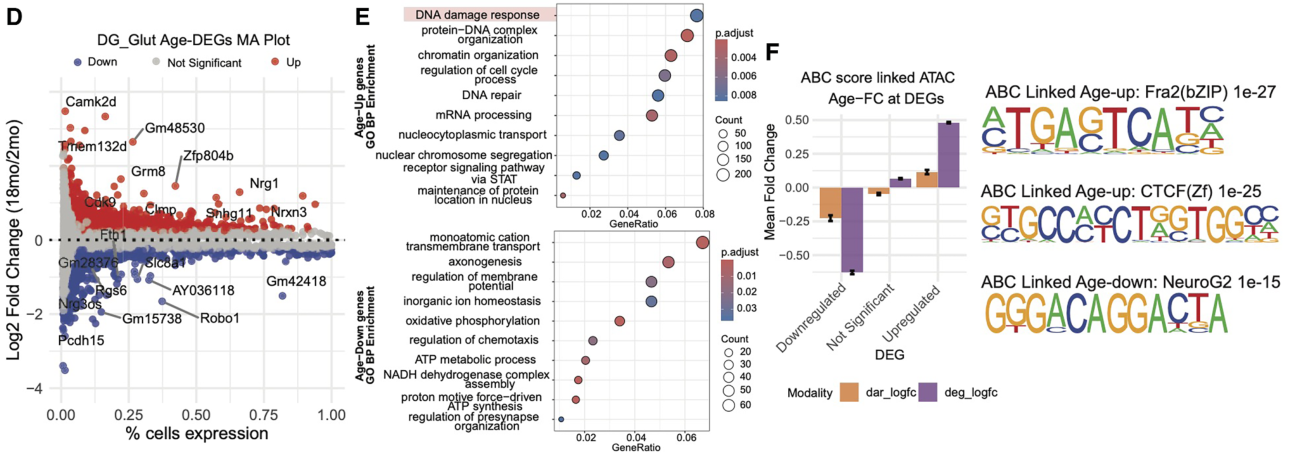
Subsequent in-depth analysis of their single-cell chromatin accessibility and transcriptomic data provided insight at three distinct levels. Firstly, the authors highlighted the progressive decline of lineage-defining transcription factor levels as a central feature of aging; their data suggested that reduced Lhx2 and Pax6 expression in progenitor cells impaired neurogenesis, reduced Sox activity in oligodendrocytes provided a mechanism underpinning age-related myelin decline, and the loss of Mef2 and NeuroG2 activity in neurons appeared to occur alongside downregulated axonogenesis and synaptic plasticity.
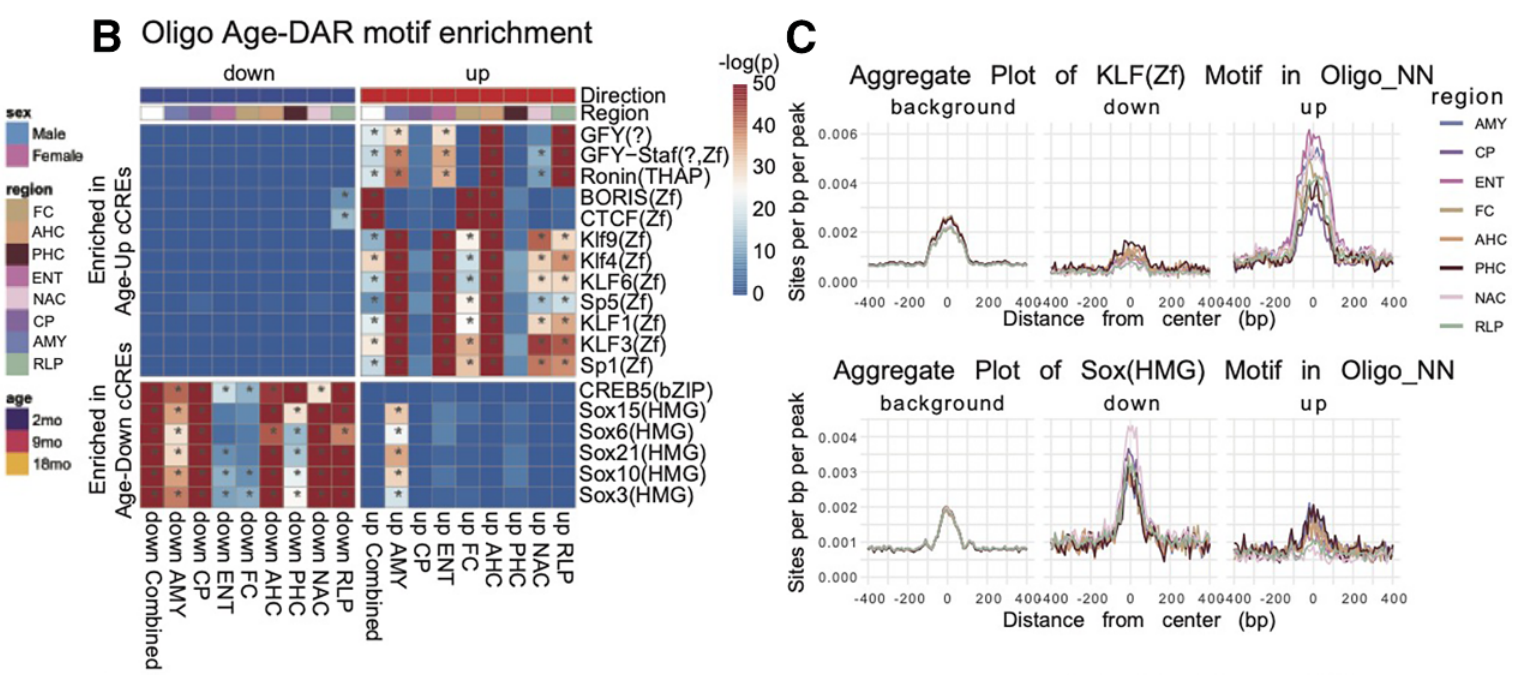
Secondly, the study revealed that aging cells exhibited increased stress-responsive transcriptional program activity as developmental transcription factor expression declined; overall, the data provided evidence for the enrichment of AP-1 motifs in neurons, KLF motifs across multiple cell types, and IRF-family motifs in microglia, linking aging to inflammatory and stress-associated regulatory states. Overall, these insights provide evidence for a progressive drift in cellular identity upon increasing age.
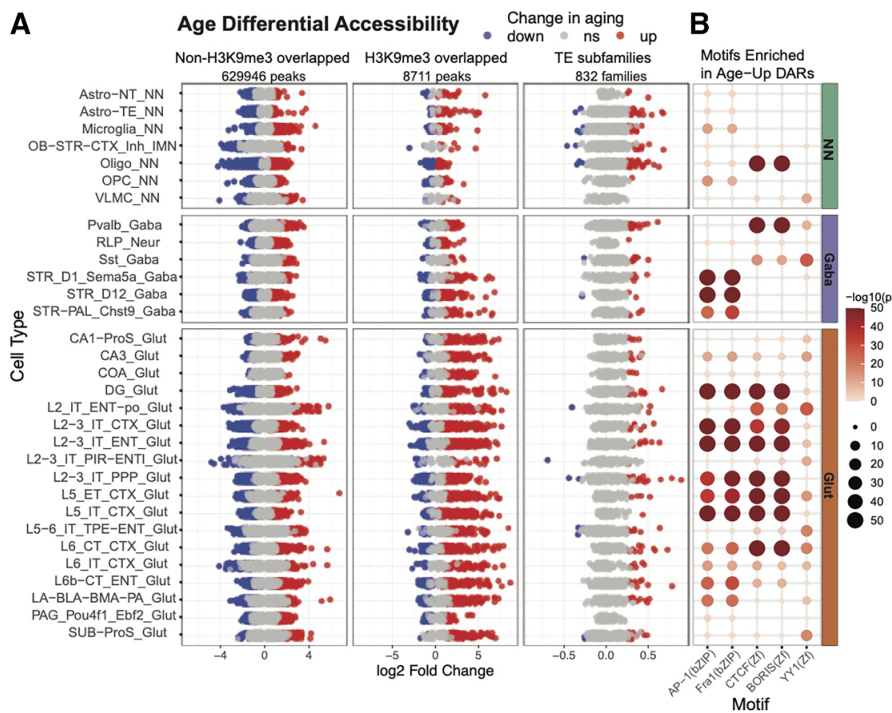
Thirdly, the authors discovered widespread heterochromatin destabilization, with associated transposable element activation and long non-coding RNA upregulation, during brain aging. In this case, the data highlighted pronounced age-associated accessibility gains at H3K9me3-marked repressive chromatin domains in glutamatergic neurons across multiple cell types and suggested the existence of sex-specific epigenetic aging mechanisms, as they observed extensive X chromosome remodeling in the female brain, which included increased accessibility at heterochromatin-associated loci and reduced accessibility at pseudoautosomal regions.
Could Paired-Tag Provide Even More Insight?
Overall, this exciting study of aging across distinct regions of the mouse brain suggests progressive epigenetic alterations and accompanying changes in gene transcription profiles that underpin normal cell identity, leaving cells in stress-responsive states that may support age-related cognitive decline and neurodegeneration. As such, these findings highlight the maintenance of chromatin states and transcription factor networks as therapeutic targets for brain preservation.
The implementation of Paired-Tag technology from Epigenome Technologies, which generates joint epigenetic and transcriptomic profiles at single-cell resolution and detects histone modifications and RNA transcripts in individual nuclei with efficiency comparable to single-nucleus RNA-seq/ChIP-seq assays, has the potential to provide deeper insight into such research aims. What more could the simultaneous single-cell analysis of histone modification and transcriptomic profiles tell us about aging in single cells isolated from regions that span the mouse brain?