PARM Predicts Promoter Activity from DNA Sequences
Predicting Promoter Activity from Underlying DNA Sequences: Still Cost- and Time-Intensive? A new study reports a promoter activity regulatory model (PARM) as an efficient means of predicting promoter activity from DNA sequences.
PARM: An Efficient New Model that Predicts Promoter Activity from DNA Sequences
The combination of deep learning techniques and datasets that aggregate transcriptional and epigenomic features across multiple distinct cell types represents one approach to constructing models that attempt to predict promoter activity from the underlying DNA sequence. However, models created using this somewhat cost- and time-intensive approach still face difficulties when aiming to infer causal links between DNA sequences and promoter activity (Barbadilla-Martinez et al. 2025). While these problems have not prevented the creation of deep-learning models that have revealed certain characteristics of human promoter regulatory "grammar" (Dudnyk et al. and Fu et al.), they still cannot predict changes in gene regulation in cell types/conditions not present in the initial training data set.
Could "massively parallel reporter assays" (MPRAs) provide an alternative, better source of training data for these models? MPRAs test DNA sequences in isolation to establish regulatory activity in a cell type of interest, providing a more straightforward way to link DNA sequences to causal roles. Encouragingly, MPRA data and deep learning have already been combined to develop powerful predictive models; however, their application to human gene promoters remains somewhat limited (Sahu et al. and Agarwal et al.).
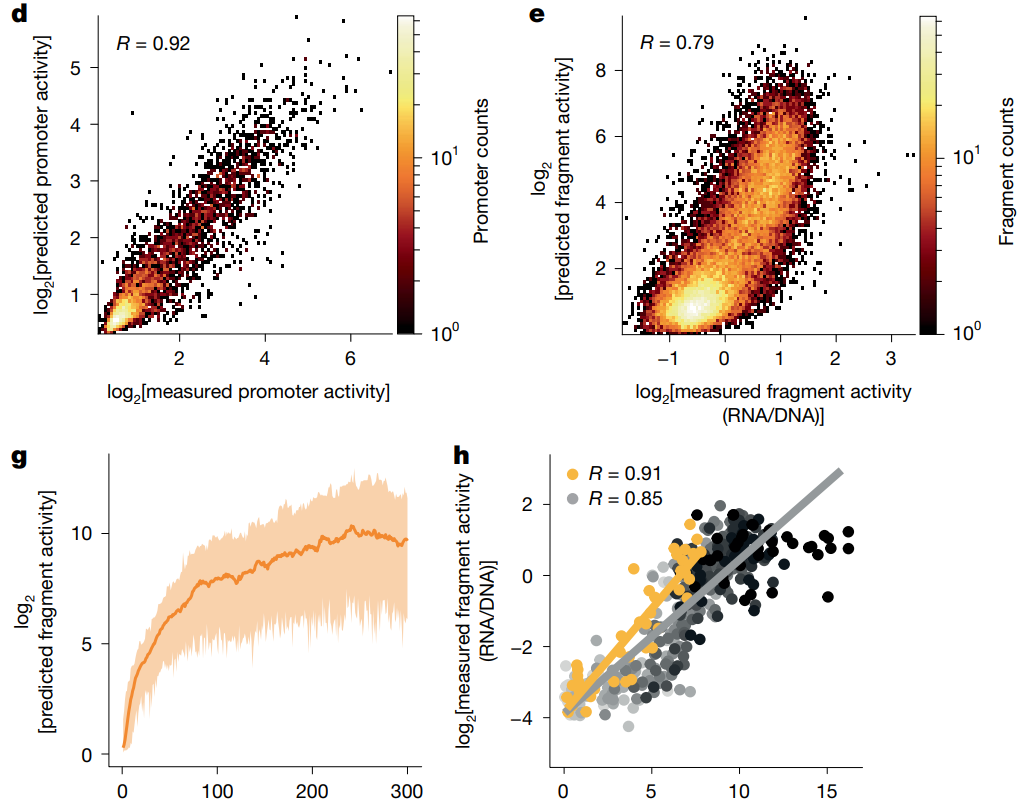
In their new study, researchers led by Jeroen de Ridder and Bas van Steensel now describe the combination of MPRAs and deep learning for the cost- and time-effective development of a promoter activity regulatory model ("PARM") for all human promoters (Barbadilla-Martínez et al. 2026). Overall, PARM can generate cell-type-/condition-specific models that predict promoter activity from DNA sequence, design strong synthetic promoters, identify transcription factor (TF) binding sites that contribute to human promoter activity, detect stimulus-induced rewiring of regulatory interactions, and confirm the complex grammar of motif-motif interactions.
PARM: MPRAs And Deep Learning as an Economical Means of Model Creation
Initial PARM construction employed training data from MPRAs previously performed by the lab, measuring the activity of ~10 million DNA fragments that overlapped a set of over 30,000 promoters in human cancer cell (van Arensbergen, et al.). This PARM successfully predicted i) the activity of randomly selected promoters excluded from the training set, individual tested promoter fragments, and genome-integrated promoters, ii) the effects of nucleotide substitutions in a promoter sequence, and iii) the impact of cis-acting expression quantitative trait loci. When compared to state-of-the-art deep learning models trained on thousands of epigenomic datasets, PARM performed as well as Enformer (Avsec et al.) but less well than Borzoi (Linder et al.), although Borzoi suffers from a significantly greater computational burden. PARM could also generate synthetic promoters with high activity and accurately pinpoint sequence elements that affect promoter activity; each promoter remained significantly dissimilar to any sequence in the human genome and carried a distinct combination of TF binding motifs, suggesting that PARM had learned aspects of human promoter regulatory grammar. Additionally, PARM identified relatively infrequent, poorly annotated functional TF motifs with unknown regulatory interactions.
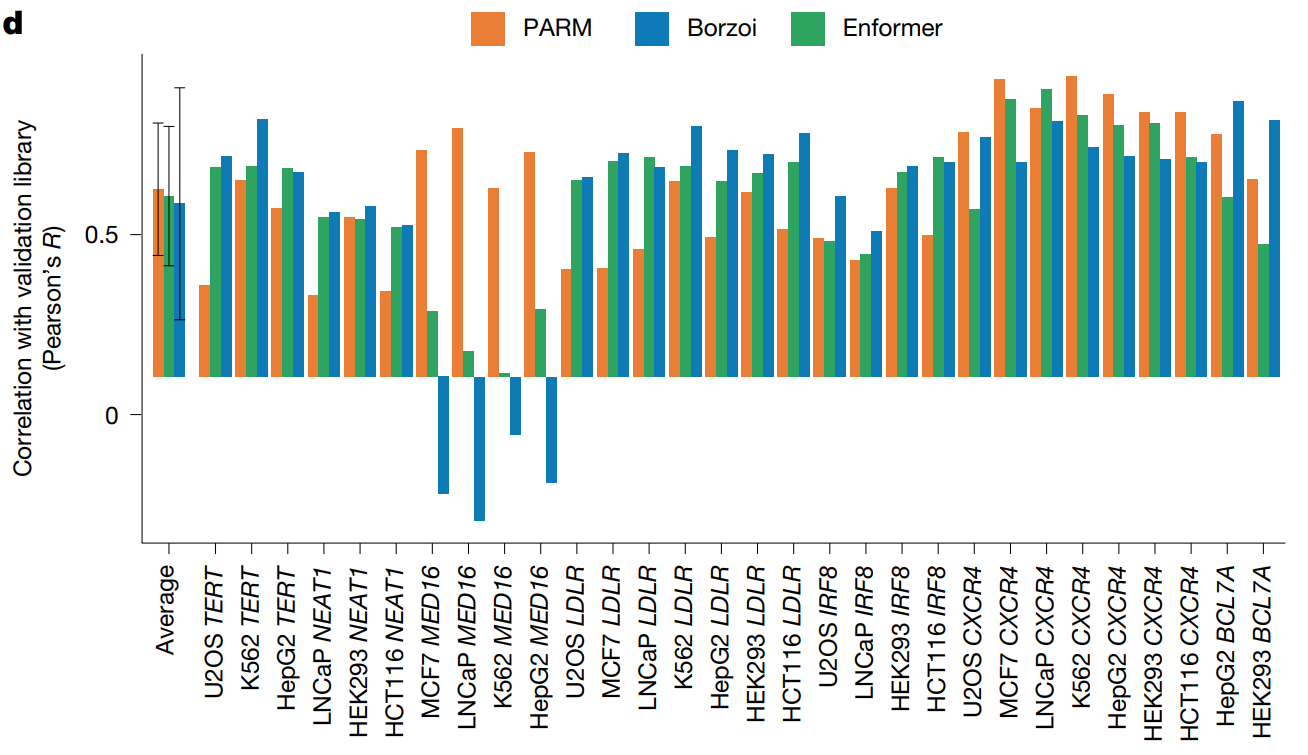
.
To reduce complexity and improve scalability, the authors next created a more "svelte" MPRA library containing only promoter-overlapping fragments to generate a new, smaller training dataset (reducing the number of cells required by 240-fold). The development of a "lightweight" PARM that compared well with the model trained on the larger datasets permitted the development of PARM models across a range of human cell lines and a patient-derived organoid culture. When employed to identify regulatory sequences, PARM (while computationally much leaner than Enformer and Borzoi) performed at a similar level and could uncover cell-type-specific promoter regulation. Overall, this new approach represented a cost-effective and versatile strategy at both experimental and computational levels.
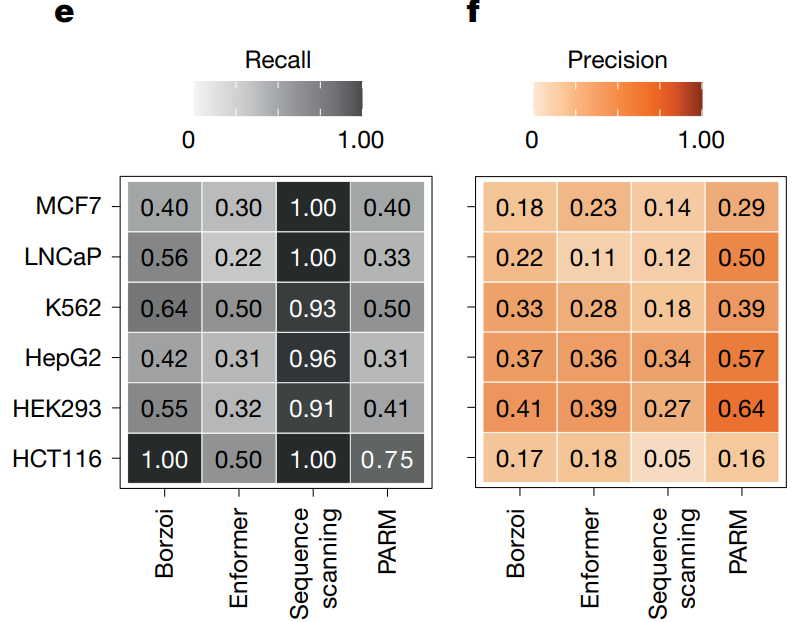
The next step in this study involved using PARM models to explore the alterations to promoter regulation that occur in response to various cell stimuli using appropriate MPRA training sets. The results highlighted the ability of PARM to i) discern specific alterations in promoter regulation associated with heat-shock responses (Himanen et al.); ii) resolve the division of labor of multiple TFs across promoters when studying TP53-proficient cells exposed to a TP53-activating drug; and iii) produce detailed information on the dynamic regulation of individual human promoters and associated TFs when treating cancer cells with a pleiotropic anti-tumor stimulus (Whalen et al.).
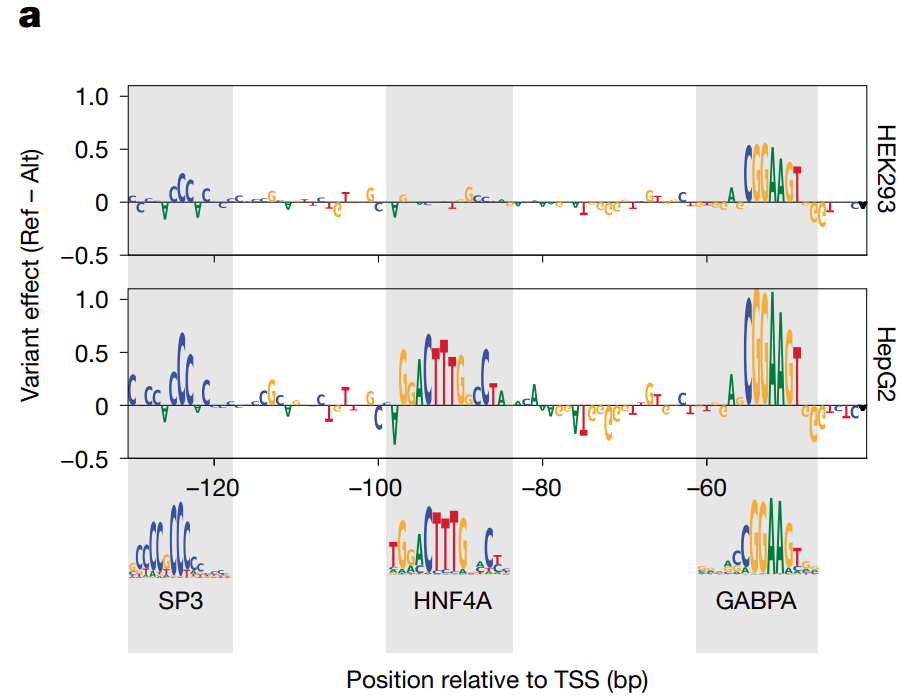
.
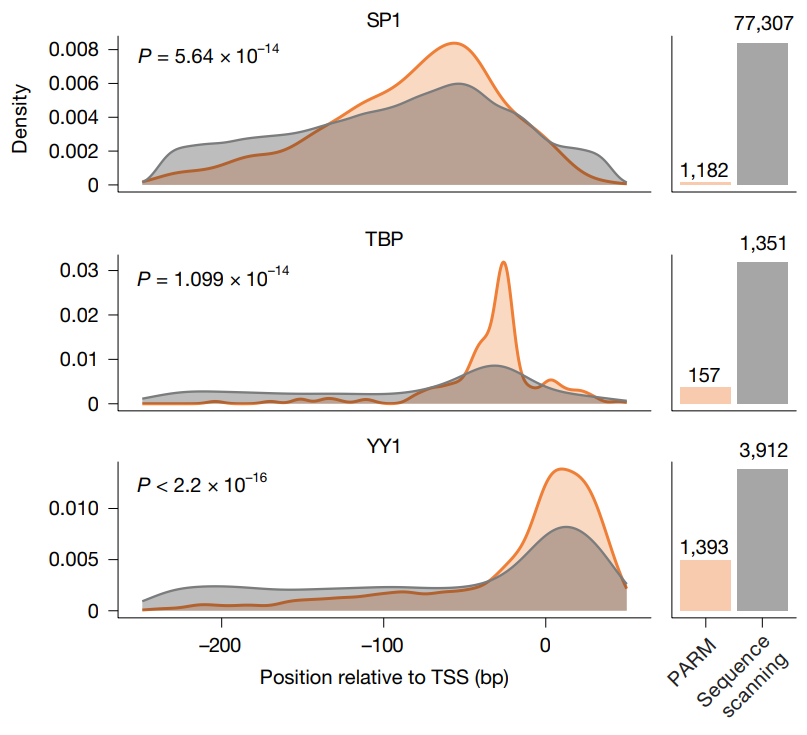
Finally, the authors studied the preferential positioning of TF binding motifs in human promoters by using PARM to predict the effects of inserting them into native promoter sequences. An initial analysis with an NRF1 motif in the FCF1 promoter revealed highly diverse position-dependent effects, while a systematic insertion of four different TF motifs throughout the over 30,000 promoter sequences revealed general trends, which included predictions that NRF1 motifs activated promoters, that NFYA and SP1 motifs induce similar impacts, and that the YY1 motif exhibited only activating effects. Overall, the data revealed that any effects depended strongly on the local sequence and the baseline promoter activity.
.
The Future of PARM
Overall, this exciting study demonstrates that combining MPRAs and deep learning constitutes an economical strategy to create models of TF regulation across human promoters in multiple cell types and in response to various stimuli. The authors highlight how the PARM captures regulatory grammar and can design artificial promoters, and, as such, represents an alternative to significantly larger modelling efforts (such as Enformer and Borzoi). Furthermore, they believe that PARM may have applications in personalized medicine, where analyses of patient-derived cell lines and organoids may provide insights into disease progression and treatment responses. Importantly, even given the hugely impressive results described, the authors believe that PARM only provides insights into the basic properties of promoters and will require further optimization of focused MPRA libraries and deep-learning methodologies, as well as the integration of data on distal enhancers, insulators, epigenetic modifications, and large-scale chromatin domains, to provide yet more insight. Of note, such advances may require the creation of hybrid models that combine MPRA data with epigenomic mapping data.
PARM is not the first sequence-only model of regulatory element activity: early assays date back to ENCODE and were subsequently used to train "CODA" to identify and design cell-specific cis-regulatory elements (Gosai and Castro et al.), and "MPRALegNet" to predict overall sequence activity (Agarwal et al.). These are efforts using quite similar, but disjoint, datasets to train these models; and the obvious next step is to refine the model weights with the expanded (joint) dataset, and to utilize the model to generate or nominate sequences with high information content to be tested. However, the ability of MPRA-derived models to predict beyond the scope of their training cells will necessarily be limited by the epigenetic factors that modulate chromatin surrounding the vector insertion. Understanding how epigenetic status interacts with regulatory sequence is therefore critical to understanding the fundamental biology of gene regulation.
Paired-Tag technology from Epigenome Technologies generates joint epigenetic and transcriptomic profiles at single-cell resolution and detects histone modifications and RNA transcripts in individual nuclei with efficiency comparable to single-nucleus RNA-seq/ChIP-seq assays. As such, Epigenome Technologies believes that profiling using Paired-Tag technology has the capacity to provide the datasets required to develop better foundational models; for more insight, check out our new Substack page, where you can read about what epigenomic foundation models reveal about biology, about epigenetic foundation models for DNA methylation, and about how single-cell epigenetics can fine-tune regulatory models such as Borzoi.