Stem Cell Fate Controlled by Novel Epigenetic Mechanism
Do Cellular Metabolism and the Nuclear Lamina Function Shape Cell Fate Through Epigenetic Mechanisms? A new study in mouse embryonic stem cells links cell metabolism, the function of the nuclear lamins, and epigenetic regulation of gene expression to cell fate.
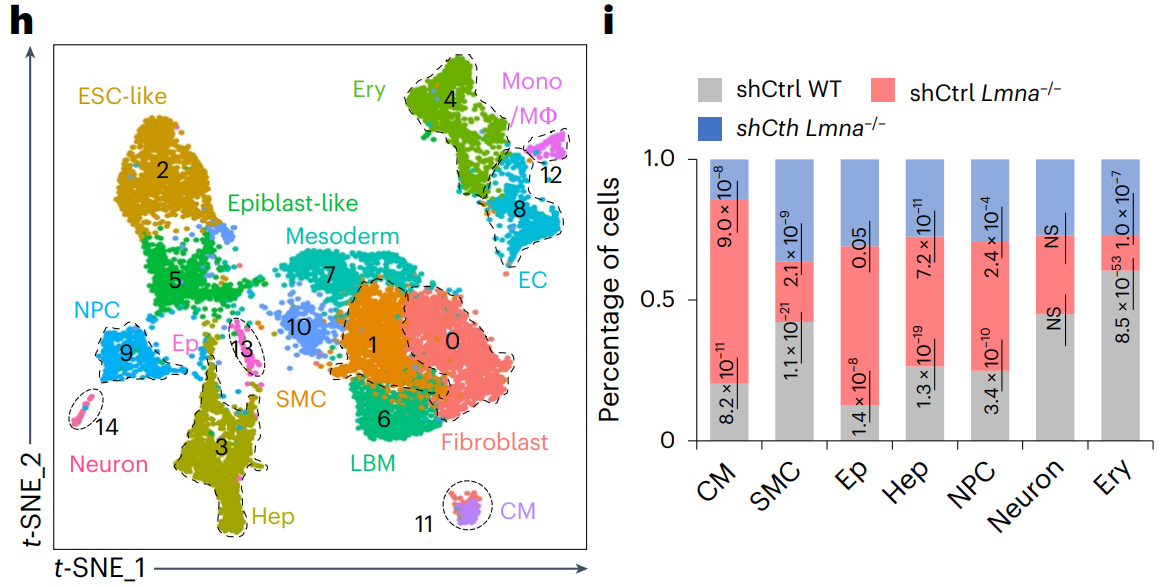
Linking Metabolism, Epigenetic Regulation, and the Nuclear Lamina to Stem Cell Fate
A vast body of research has linked the self-renewal and differentiation of pluripotent stem cells to specific transcriptional, epigenetic, and metabolic profiles; furthermore, the distinct types of pluripotency observed in vitro - including "naïve" and "primed" pluripotency (Nichols & Smith and Weinberger et al.) - also differ regarding transcriptional profiles, epigenetic landscapes and metabolic activity (Wu et al. and Sperber et al.). Importantly, the known relationship between cell metabolism and epigenetic/ transcriptional regulation arises, in part, because certain metabolites serve as cofactors for epigenetic modifiers. For example, the bioavailability of cysteine - a major intermediate product of cellular amino-acid metabolism - controls the levels of acetyl-CoA and S-adenosylmethionine, which regulate histone acetylation and methylation, respectively.
Interestingly, genome-nuclear lamina interactions also determine cell-fate decisions in pluripotent stem cells; in short, nuclear lamins can anchor lineage-specific genes to the nuclear periphery, keeping them in a silent state. Overall, this mechanism can ensure precise lineage specification and tissue morphogenesis by controlling the chromatin composition of these gene loci and, hence, their expression. As such, the majority of genes associated with the nuclear lamina remain transcriptionally inactive and marked by repressive histone modifications (Reddy et al. and Wen et al.). Studies have reported the dramatic downregulation of lamin A/C expression as pluripotent stem cells transit from a naïve to a primed pluripotent state (Wang et al.) and the aberrant activation of cell-fate genes and abnormal cell fate in response to lamin A/C loss or mutation (See the original article for extensive references).
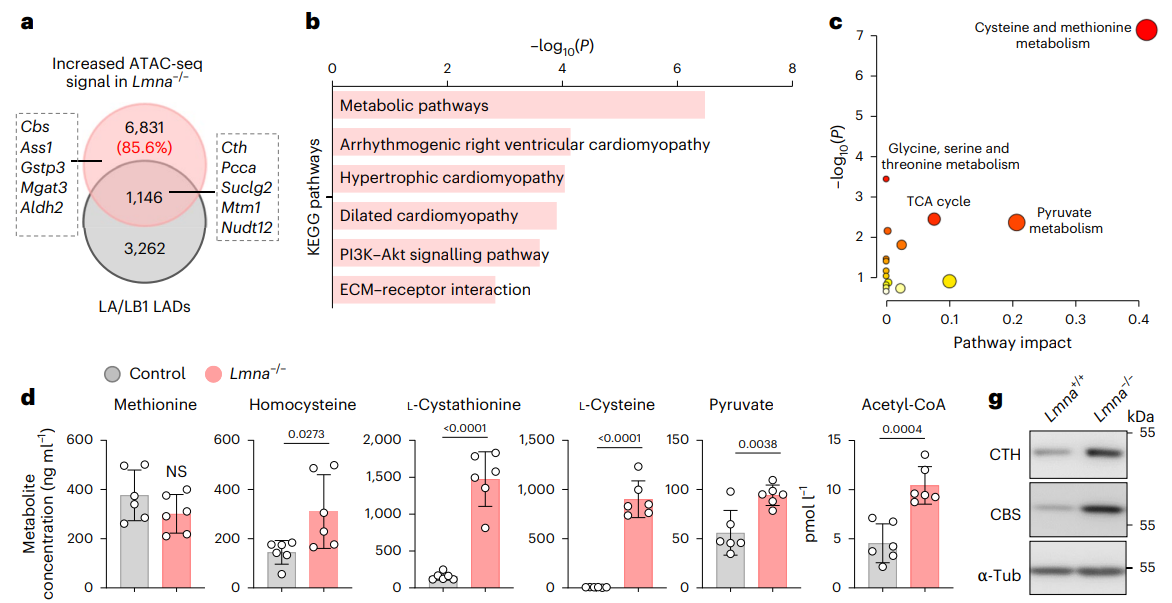
Overall, changes in cellular metabolism and the nuclear lamina shape cell fate through epigenetic mechanisms; is there a link between these two seemingly disparate fields? Addressing this question directly, researchers led by Yinuo Wang, Sofia-Iris Bibli, and Gergana Dobreva now establish that lamin A/C regulates cysteine metabolism in mouse embryonic stem cells and, as such, represents a critical regulator of cell fate and longevity. Overall, Wang and Shi (et al.) hope that their new study in Nature Metabolism will advance our understanding of the relationships present between cell metabolism, epigenetics, genome-nuclear lamina interactions, and cell fate, and support the development of novel therapeutics for lamin-associated disorders such as Hutchinson-Gilford progeria syndrome.
Paired-Tag technology from Epigenome Technologies generates joint epigenetic and transcriptomic profiles at single-cell resolution and detects histone modifications and RNA transcripts in nuclei with efficiency comparable to single-nucleus RNA-seq/ChIP-seq assays. Could integrating Paired-Tag have helped explore the links between cell metabolism, epigenetics, genome-nuclear lamina interactions, and cell fate at the single-cell level, and provide deeper insight?
Lamin A/C, Cysteine Catabolic Flux, and Epigenetic Modifying Enzymes Control Embryonic Stem Cell Fate
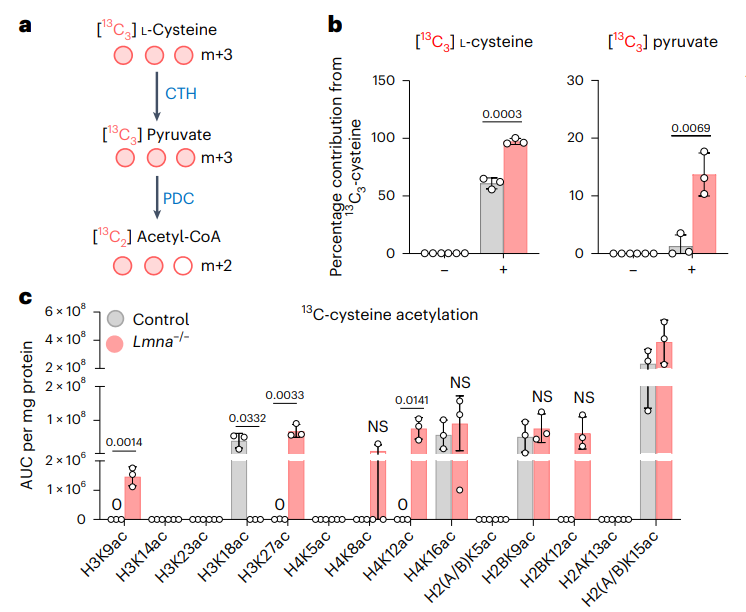
Previous research by the same authors had revealed that lamin A/C helps to control 3D chromatin organization in mouse embryonic stem cells, ensuring naïve pluripotency and aiding proper cell-fate choices during differentiation (Wang et al.). In this new study, the authors discovered that the high level of Lamin A/C observed in naïve pluripotent stem cells suppressed the expression of metabolic and differentiation-associated genes, and that the transition to a primed pluripotent state occurred alongside a downregulation of Lamin A/C levels. The authors knew that Lamin A/C could restrict transcription factor binding via the recruitment of repressive epigenetic complexes (Wang & Dobreva); here, they discovered that reduced lamin A/C levels prompted a reduction in the levels of the repressive H3K27me3 histone modification at the promoters of the cysteine-generating and -metabolizing enzymes (cystathionine γ-lyase and cystathionine β-synthase) via reduced polycomb repressive complex 2 (PRC2) activity, which, in turn, allowed the binding of the SP1 regulatory transcription factor and elevated gene expression (Maclean et al. and Yang et al.). The subsequent increase in de novo cysteine synthesis then supported elevated levels of H3 acetylation at K9 and K27 through greater acetyl-CoA abundance, thereby facilitating epigenome remodelling events crucial to stem cell differentiation and germ layer formation.
Subsequent studies in mouse embryonic stem cells expressing a dominant gain-of-function mutation in the Lmna gene (which encodes Lamin A/C) that prompts premature ageing (Eriksson et al.) revealed a decrease in cysteine availability and catabolic flux, the rerouting of cysteine catabolic flux due to reduced cystathionine β-synthase levels, and the increased recruitment of the repressive epigenetic complex SUV39H1 to chromatin and an increase in the levels of H3K9me3. Interestingly, cystathionine β-synthase re-expression restored many of the observed deficiencies in Lmna mutant embryonic stem cells associated with aging, including restored DNA repair, reduced mitochondrial reactive oxygen species production, reduced senescence-associated secretory phenotype gene expression, and prevented senescence, which may arise from cystathionine β-synthase limiting the availability of S-adenosylmethionine, thereby attenuating SUV39H1-dependent H3K9me3 deposition.
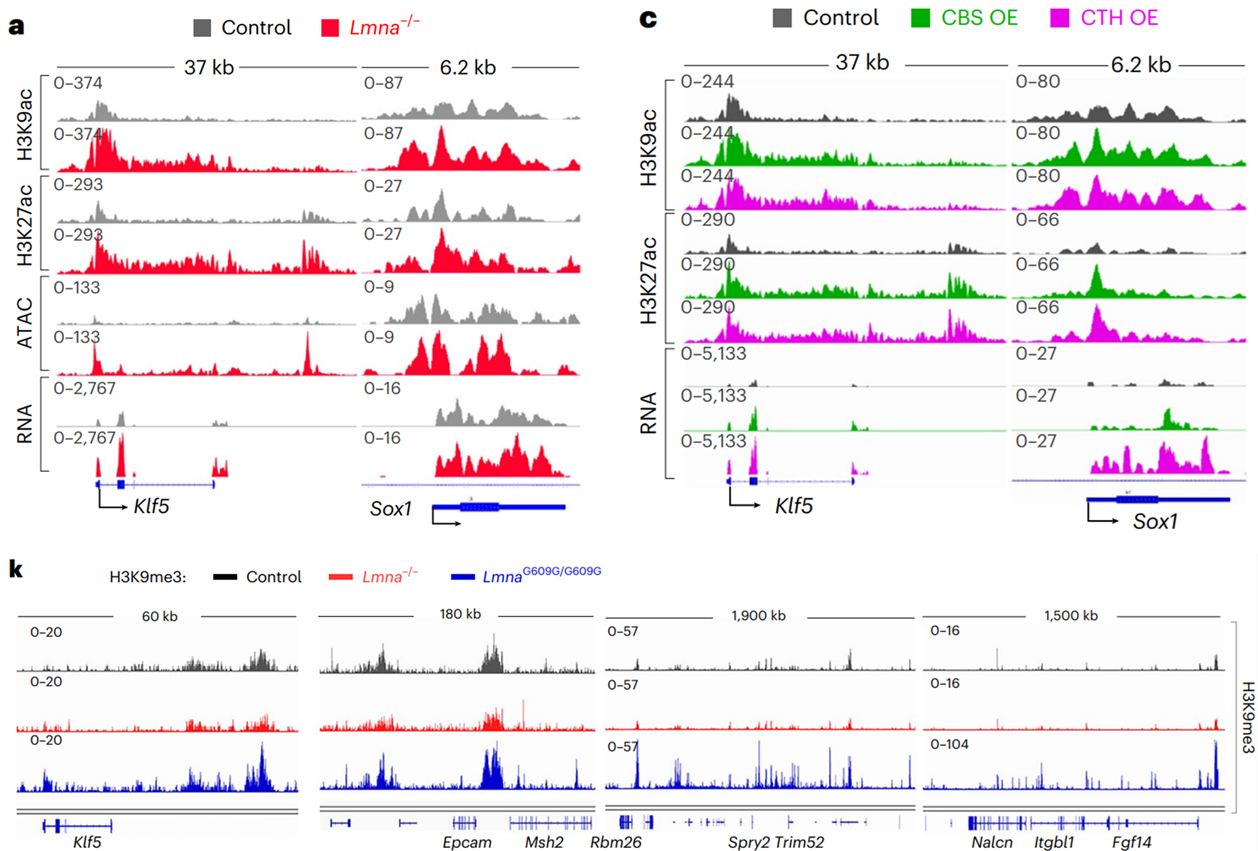
Towards The Development of Novel Therapeutic Strategies for Lamin-Associated Disorders
This exciting study - revealing that abnormal cysteine metabolism during early embryogenesis alters cell fate, function, and longevity - may suggest novel therapeutic strategies for lamin-associated disorders; for example, could dietary interventions hold promise in disease prevention by counteracting perturbations in genes encoding epigenetic modifiers? Such advances may require renewed efforts to understand the dynamic interface of epigenetic alterations and metabolism in development, homeostasis, and healthy ageing.
The implementation of Paired-Tag technology from Epigenome Technologies, which generates joint epigenetic and transcriptomic profiles at single-cell resolution and detects histone modifications and RNA transcripts in individual nuclei with efficiency comparable to single-nucleus RNA-seq/ChIP-seq assays, has the potential to provide deeper insight into such research aims. What more could the simultaneous single-cell analysis of histone modification and transcriptomic profiles tell us about the crossroads of cell metabolism, epigenetics, genome-nuclear lamina interactions, and cell fate in mouse embryonic stem cells?